Best practices, a computer package, and hands-on tutorials to guide the suggested application of a powerful emerging technique for prediction of plant traits from hyperspectral data.
The Science
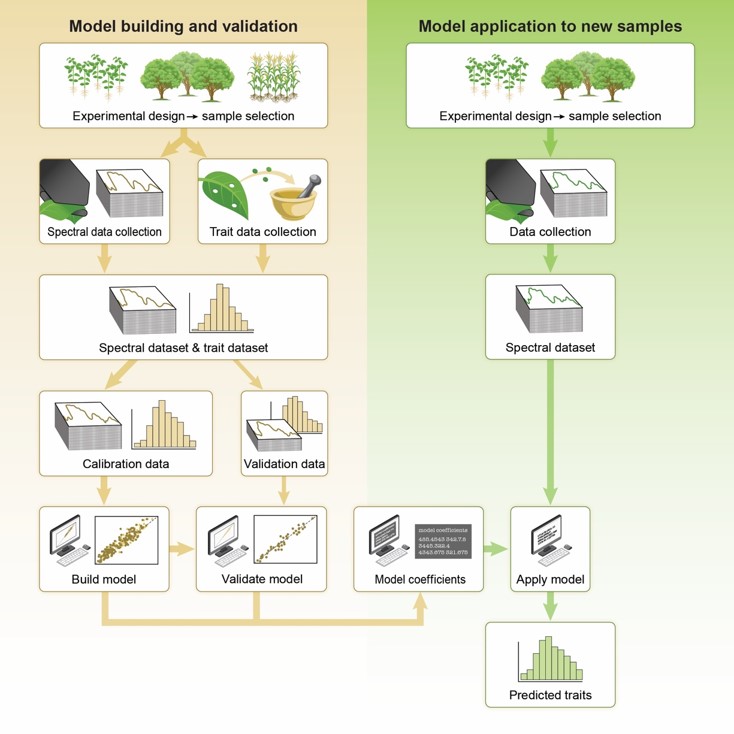
The estimation of leaf traits, such as leaf nitrogen, from hyperspectral reflectance data enables rapid, high-throughput, non-destructive characterization of leaf function and plant phenotyping with applications in ecosystem characterization and monitoring. Partial least squares regression (PLSR) is a popular method for the prediction of a wide range of nutritional and structural leaf traits using hyperspectral data. However, there has not been a standard approach for developing and reporting these models and results. This has led to challenges in the wider application of the technique. We developed a detailed description of use of PLSR to predict leaf traits with spectra and our recommended best practices across all steps of the process, from experimental design, data collection, PLSR model building, model application and reporting of results. With the article, we also provide hands-on tutorials to assist users to in understanding the PLSR modeling and application of our best practices with their own data.
The Impact
Plant scientists require detailed and extensive information on the concentration and distribution of physiological and structural leaf properties to study vegetation responses to environmental change, monitor plant health, and facilitate the rapid screening of different plant phenotypes. Traditional approaches to measure these traits directly are expensive and logistically challenging. Here we summarize and illustrate an alternative spectroscopic approach for the rapid, accurate and non-destructive estimation of traits using remote sensing data.
Summary
Plant physiologists and ecologists regularly measure leaf functional traits, including leaf nitrogen or photosynthetic rate, across a range of leaves, plants, species, or environments. These direct measurements, while very accurate for characterizing leaf structure and function, are typically slow, expensive, and can be logistically challenging. In addition, many ecological or phenotyping studies require a large number of samples, which can be impractical with traditional methods. On the other hand, remote sensing methods have been shown to be effective for the rapid estimation of many of key leaf traits, however the inconsistent usage of the methods have led to challenges in the wider application across the plant sciences. To address this challenge, and to help standardize the approach across studies to facilitate wider adoption, we provide a detailed summary of the spectral method of leaf trait estimation. We also provide clear examples and tutorials to illustrate how to use the approach, and we discuss a range of suggested “best-practices”. Importantly, we also highlight how the same approach can be scaled up to estimate vegetation traits across landscapes using non-contact remote sensing data.
Contact (BER): Daniel Stover, SC-23.1 (Daniel.Stover@science.doe.gov)
Science Contact: Shawn P. Serbin, Brookhaven National Laboratory, sserbin@bnl.gov (+1 631-344-3165)
Funding
This work was supported by the Next-Generation Ecosystem Experiments (NGEE Arctic and NGEE Tropics) projects that are supported by the Office of Biological and Environmental Research in the Department of Energy, Office of Science, and through the United States Department of Energy contract No. DE-SC0012704 to Brookhaven National Laboratory.
Publications
Burnett, A. C., J. Anderson, K. J. Davidson, K. S. Ely, J. Lamour, Q. Li, B. D. Morrison, D. Yang, A. Rogers, and S. P. Serbin. 2021. “A best-practice guide to predicting plant traits from leaf-level hyperspectral data using partial least squares regression”. Journal of Experimental Botany. [DOI: https://doi.org/10.1093/jxb/erab295]
Related Links
Article URL: https://doi.org/10.1093/jxb/erab295
Companion computer code: https://zenodo.org/record/4730995
Brookhaven National Laboratory “Of Leaves and Light” article on the use of spectroscopy to estimate leaf functional traits https://www.bnl.gov/newsroom/news.php?a=213279
Open-source book chapter, “Scaling Functional Traits from Leaves to Canopies”: https://link.springer.com/chapter/10.1007/978-3-030-33157-3_3